|
Today, businesses of all sizes use barcode software and hardware to create a technological environment that improves accuracy, efficiency, workflows, and profitability in their organization. Benefits are realized throughout the company and the capabilities gained are available without leaving QuickBooks. A barcode system significantly improves inventory management, warehousing and order fulfillment and most businesses find the return on investment can be realized within a few months, while the benefits are ongoing for years to come. The cost of errors for all businessesAll businesses, especially distribution businesses, face the high costs of errors. Regardless of industry, barcoding significantly improves business operations and customer satisfaction. Transform how your entire business operates with QuickBooks barcode software and technologyHow can barcodes improve your business?Accuracy. A Barcode system improves accuracy and enables processes to be implemented that reduce human errors and improve inventory management discipline.Speed up.
Scanning barcodes is significantly faster and easier than documenting and then typing information.Cost savings. Accuracy improvements & time savings, in addition to the reduction of fixing costly mistakes, can result in considerable cost savings across an organization.Operational efficiency.
Operational bottlenecks and productivity issues can more easily be identified and remedied using barcoding in workflow management.Customer service. Accurate inventory and stock levels reduce out of stocks, improving customer satisfaction in the process and reducing time spent on resolving customer problems.Decision-making. Rapid, accurate data collection enables access to real-time business intelligence across all areas of your company.Tracking & tracing. Lot numbers can be recognized when received, shipped or both, enabling compliance with regulatory requirements for specific industries.Acctivate QuickBooks barcoding software user, Ugly Mug CoffeeHow do you get started using barcodes in your business?By first gaining an understanding of the components of a barcode system, it is then possible to determine your needs and the applications that could be implemented in your business.If you are new to barcoding, you may want to start with our article, “,” which is a simple overview of barcodes and what they can and cannot do.
The components of a barcoding systemBarcode management is the process of creating unique barcodes to identify individual items.The system includes the software and hardware required to create and scan barcoded labels & documents and associate the barcodes with the correct items in the inventory management system. Barcode software, such as Acctivate, is the key driver of the system.
Species substitution is a form of seafood fraud for the purpose of economic gain. DNA barcoding utilizes species-specific DNA sequence information for specimen identification. Previous work has established the usability of short DNA sequences—mini-barcodes—for identification of specimens harboring degraded DNA. This study aims at establishing a DNA mini-barcoding system for all fish species commonly used in processed fish products in North America. Six mini-barcode primer pairs targeting short (127–314 bp) fragments of the cytochrome c oxidase I ( CO1) DNA barcode region were developed by examining over 8,000 DNA barcodes from species in the U.S. Food and Drug Administration (FDA) Seafood List. The mini-barcode primer pairs were then tested against 44 processed fish products representing a range of species and product types.
Of the 44 products, 41 (93.2%) could be identified at the species or genus level. The greatest mini-barcoding success rate found with an individual primer pair was 88.6% compared to 20.5% success rate achieved by the full-length DNA barcode primers. Overall, this study presents a mini-barcoding system that can be used to identify a wide range of fish species in commercial products and may be utilized in high throughput DNA sequencing for authentication of heavily processed fish products. Food fraud from species substitution is an emerging risk given the increasingly global food supply chain and potential food safety issues. Economic food fraud is committed when food is deliberately placed on the market, for financial gain, with the intention of deceiving the consumer.
As a result of increased demand and the globalization of the seafood supply, more fish species are being encountered in the market. In fact, the Seafood List from the U.S. Food and Drug Administration (FDA) contains more than 1,700 acceptable market names that can be used to label seafood in interstate commerce in the U.S.
Subsequently, the need for accurately labelled food products and full disclosure of product composition has become more critical. One difficulty in this is the authentication process of different seafood products through examination of the physical appearance of specimens. In their whole or unprocessed form, these species can generally be identified based on morphological indicators; however, over half of the fresh/frozen finfish imported into North America is processed from its original form into products such as fillets and steaks, blocks, and fish sticks. Moreover, species identification by morphological indicators requires a certain level of expertise to distinguish between closely related species. Unfortunately, most consumers are unable to detect cases of mislabelling or fraud given that recognizable external morphological features are typically removed when the fish is processed.To audit and prevent species fraud on the commercial market, a number of molecular methods have been developed, including use of a unique protein or DNA profiles found in different species. DNA barcoding provides a rapid, cost-effective method for accurate identification at the species-level through comparative analysis of sequence variation in a short, standardized fragment of the genome.
The designated DNA barcode for animal species identification is a 650-bp fragment of the mitochondrial gene coding for cytochrome c oxidase 1 ( COI). A number of studies have shown the applicability of DNA barcoding for accurate identification of a wide range of fish species. Recently, DNA barcoding has been employed as a species identification tool for food authentication and safety concerns, including incorrect product labelling, ingredient substitutions or food contamination, as well as for regulatory use. DNA-based methods can also be used to monitor illegal trading involving protected or endangered species, or to identify the species origin of commercially processed food. However, some of the processing and preservation methods used with seafood products are not conducive to DNA barcoding with the full-length target gene region. DNA degradation has been recognized as a considerable limitation in DNA-based analyses of these samples, and PCR amplification of full-length (i.e.650 bp) barcodes from moderately or highly processed samples is significantly challenging.
In addition, processed seafood products often contain multiple additives, preservatives, and flavors that may affect the quantity and quality of DNA extracted from these products. Alternatively, a mini-barcoding approach, which focuses the analysis on shorter DNA fragments (e.g., 100–200 bp) within the full-length barcode, has been shown to be effective in obtaining DNA sequence information from specimens containing degraded DNA. The sequencing information generated from a small (≥100 bp) mini-barcode fragment of COI within the full-length DNA barcode region can provide the information required for identification of individual species with more than 90% species resolution. However, extensive mini-barcode primer development and testing specifically for use with commercially processed fish species has not been carried out.Here, we designed and optimized multiple primer sets to amplify mini-barcodes within the COI barcode region. These mini-barcode primer sets were then used to identify species in a variety of commercially processed fish products obtained in the United States. Sample collectionA total of 96 authenticated fish muscle tissue samples were obtained, representing 88 different species.
The fish tissue samples were supplied by the FDA-Center for Food Safety and Applied Nutrition. These samples are from the FDA’s Reference Standard Sequence Library for Seafood Identification and all are linked to authenticated, vouchered specimens. These samples were used for construction of a DNA barcode library, as described below. Also they were used for optimization of mini-barcode primers designed in this study. For analysis of mini-barcode primers with commercial products, a total of 44 heavily processed seafood products representing a variety of species and product types were purchased in the United States in May 2012 from online retail sources.
Subsamples were collected from each product using sterile forceps and scalpels and stored in 1.5 ml microcentrifuge tubes at −70 °C. These subsamples were shipped to the Biodiversity Institute of Ontario at the University of Guelph for DNA extraction and sequencing. DNA extractionFor each authenticated or commercial sample, one gram of tissue/product was divided into 10 MP lysing matrix tubes “A” (100 mg each) and homogenized using an MP FastPrep-24 Instrument (MP Biomedicals Inc.) at speed 6 for 40 S. Total DNA of this homogenized slurry was extracted using the Nucleospin tissue kit (Macherey-Nagel Inc.) following the manufacturer’s instructions and eluted in 50 μl of molecular biology grade water. DNA barcode library constructionThe COI standard barcoding region (652 bp) was amplified for each of the 96 authenticated samples using a pair of newly designed degenerate fish primers as well as a primer cocktail previously described.
Each amplification reaction contained 2 μl DNA template, 17.5 μl molecular biology grade water, 2.5 μl 10X reaction buffer, 1 μl MgCl2 (50 μM), 0.5 μl dNTPs mix (10 mM), 0.5 μl forward primer (10 μM), 0.5 μl reverse primer (10 μM), and 0.5 μl Invitrogen’s Platinum Taq polymerase (5 U/μl) in a total volume of 25 μl. The PCR conditions were initiated with a heated lid at 95 °C for 5 min, followed by a total of 35 cycles of 94 °C for 40 S, 51 °C for 1 min, and 72 °C for 30 S, and a final extension at 72 °C for 5 min, and hold at 4 °C. PCR reactions were carried out using Mastercycler ep gradient S (Eppendorf, Mississauga, ON, Canada) thermal cyclers. PCR success was verified by 1.5% agarose gel electrophoresis.
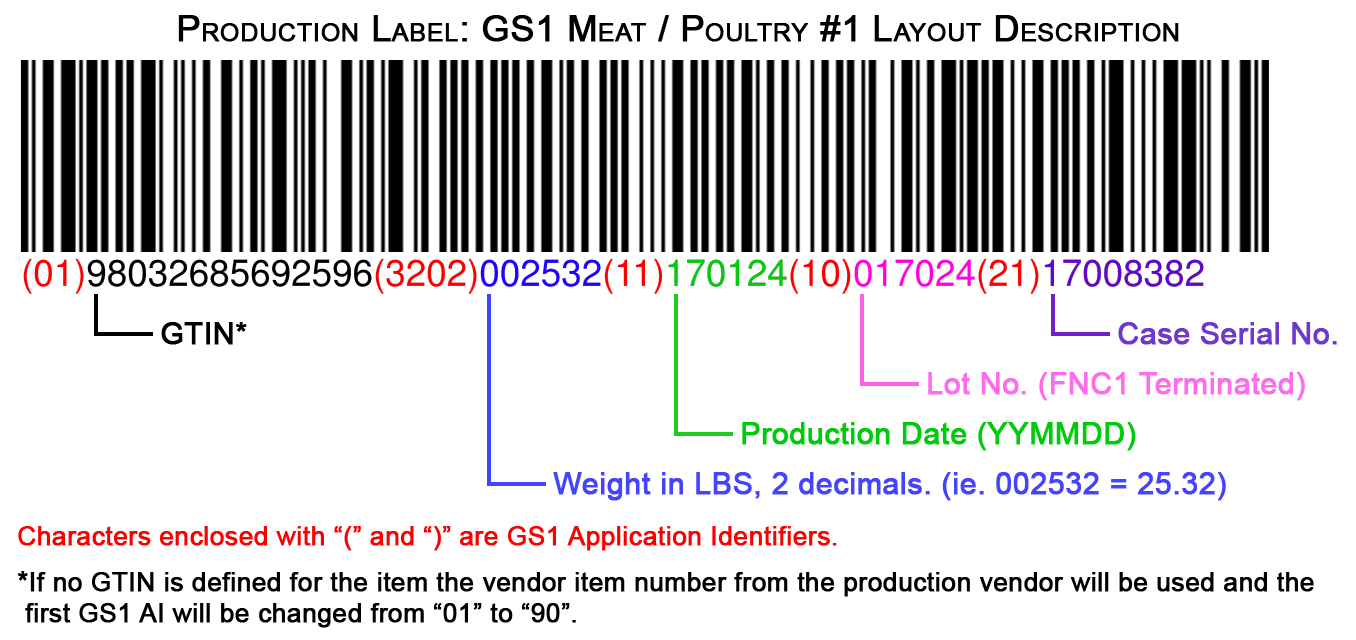
Barcode Inventory System
A DNA template negative control reaction was included in all experiments to test for contamination. Two microliters of each amplicon were subsequently used directly for bi-directional Sanger sequencing with the M13 primers described in using Applied Biosystems’s BigDye Terminator chemistry V3.1 (Foster City, CA, USA). Sequencing reactions were cleaned using EdgeBio’s AutoDTR96 (Gaithersburg, MD, USA) and visualized on an ABI 3730xl sequencer (Applied Biosystems). Sequence editing and contig assembly were carried out using CodonCode Aligner v 3.7.1.1 (CodonCode Corp., Dedham, MA, USA). Identification of the tested samples was conducted using BLAST in GenBank and a local barcode library for selected taxa with a minimum BLAST cut off of 98% identity for a top match. The accession numbers of the generated sequences are available in the.

PCR primer design and in silico testingA total of 8845 fish COI barcodes were downloaded from GenBank (n = 1894) and the Barcode of Life Database (BOLD; n = 6951) using the FDA Seafood List as a guide for the target species. All sequences were aligned and multiple copies of identical sequences were removed. Degenerate nucleotides and inosine were used to manually design a fish COI primer set to amplify 652 bp—the standard barcoding region—within a wide range of fish species.The newly designed COI primer set was used to amplify the full-length DNA barcode in the 96 authenticated samples from the FDA.
For comparison, a previously designed primer cocktail was also used to amplify these samples. The COI sequences generated from the authenticated samples, along with the unique COI sequences downloaded from GenBank and BOLD, were then used to design multiple mini-barcode primer sets to amplify partial fragments within the standard COI barcoding region. The primers were picked according to the availability of highly-conserved priming sites in a wide range of species with consideration of the primer stability in PCR reactions as well as the physical and structural properties of oligos (e.g., annealing temperature, G+C percentage, hairpin formation, and self- and hetero-dimer formation). In silico analysis was also carried out using UCLUST and MEGA V5.2.2 on the newly designed mini-barcode primers to assess the potential for the amplification targets to differentiate fish species at the 98% and 100% levels ( and Table S2).
The analysis included full-length DNA barcodes representing 200 species and 124 genera obtained from the FDA’s Reference Standard Sequence Library for Seafood Identification. M13 forward and reverse tails were attached to the forward and reverse barcoding primers, respectively, to facilitate high-throughput sequencing.
The Integrated DNA Technologies (IDT) analysis tool was used to evaluate all the mentioned parameters. Six mini-barcode primer sets were selected for further testing with commercial samples.
Mini-barcoding PCR Optimization StrategyThe amplification conditions for all the primer sets were tested using a gradient PCR approach at a wide range of annealing temperatures (43–60 °C). The composition of the amplification reactions, the PCR amplification conditions (except the annealing temperature), and the sequencing conditions were exactly the same as those used previously for amplification and sequencing of the full-length barcode. The optimal annealing temperature of each primer set was determined based on the results of gel electrophoresis of temperature gradient PCR products and is listed in. The mini-barcode amplification and sequencing steps were carried out on DNA from the 44 commercial fish products using each of the designed six sets of mini-barcode primers. Reagent blanks and a non-template PCR control were included in all PCR and sequencing experiments. Sequence editing and contig assembly of the generated barcodes were carried out as described for the full-length barcodes using CodonCode Aligner v 3.7.1.1 (CodonCode Corp., Dedham, MA, USA).
Species identification for each sample was conducted using BLAST against GenBank and a local barcode library for selected taxa with a minimum BLAST cut off of 98% identity for a top match. These results were verified by neighbour-joining analysis and subsequent evaluation of the grouping of specimens tested as compared to database sequences. Full-length DNA barcodes (652 bp) could be recovered using the newly-designed Fish primers (Fish UnivF and Fish UnivR) in 86 out of 88 of fish species (93 out of 96 specimens) within the authenticated fish muscle tissue sample collection obtained from the FDA.
Both peak intensities and sequencing qualities of the generated barcodes were compared to the sequences generated with the previously used primer cocktail. The new full-length barcode fish primer set showed slightly higher success rate (97.7%) among the wide variety of the tested species compared to a success rate of 95.5% for those species sequenced with the previously developed primer cocktail.Regarding the commercial fish products, the tested products included a wide range of processed products packed as cans, tins, retort pouches, jars, or tubes. These samples were all shelf-stable, preserved products that had experienced different levels of processing, for instance, smoking, salting, etc., and they also contained multiple additives, preservatives, and flavors. These traits may negatively impact the quantity and quality of DNA extracted from these samples, which can decrease subsequent DNA barcode recovery. The standard COI barcoding of the 44 tested fish processed products was achieved in only 9 products (20.5%) using both the newly designed universal fish primer set and the previously used fish primer cocktail.
These full-length barcodes were generated from a variety of samples with different levels of processing and with a variety of additives. The major cause of full-barcode failure was most likely due to the degradation of the DNA extracted from these samples as a result of different levels of processing and the presence of multiple additives. Samples showed low amplicon yields in gel electrophoresis and poor quality sequences with co-amplification of multiple non-targets (results are not shown).Previous work has shown the applicability of a mini-barcoding approach in different groups of organisms. Furthermore, it has been shown that the sequencing information of any 100 bases or more within the standard COI barcoding region can distinguish up to 91–94% of species in different taxonomic groups. Here, we developed primers to amplify 6 mini-barcodes for commercial fish species, based on species described in the FDA Seafood List.
Barcode System For Meat Machine
The target fragment size ranges between 127 bp and 314 bp. When compared in silico using DNA barcodes from authenticated FDA fish specimens representing 124 genera and 200 species, the mini-barcode amplification targets showed high levels of differentiation at both the species and genus levels ( and Table S2). Overall, primer sets SH-B, SH-D, SH-E, and SH-F showed the greatest ability to resolve sequences at the genus and species levels. All four of these primer sets showed high potential for use in fish species identification, with resolution at the species level for 98–100% of the species analyzed at the 100% sequence identity level and 98–99% of the species analyzed at the 99% sequence identity level. Demonstrates the amplification regions of the designed mini-barcode primers within the full-length COI DNA barcode. These mini-barcodes target 5 ′ (SH-A, SH-C, SH-E) and 3 ′ (SH-B, SH-F) regions of the standard DNA barcode as well as the middle region (SH-E, SH-D, SH-F).
Hence, their combination can maximize recovery of sequence information from across the full-length DNA barcode and should provide sufficient sequence information for species identification. In support of this, when the results of the in silico taxonomic analyses for all six mini-barcode primer sets were combined, species-level resolution was possible in 100% of sequences analyzed (Table S2).
However, it is important to note that this analysis was restricted to sequences from authenticated specimens representing 200 species of commercial fish. Incorporation of sequences from a wider number of fish species may lead to less definitive results and may require slight primer modifications. Out of the 44 processed fish products tested with the mini-barcode primer sets, 41 products (93.2%) could be mini-barcoded with at least one primer set. Three samples (RB-194, RB-1104, and RB-2114) were negative in both standard barcoding and mini-barcoding with all primer sets.
These samples were all labelled as tuna products (2 retort pouches of light tuna and 1 can of albacore) and contained a variety of additives. Although they showed some amplification success with the mini-barcoding primers, all generated amplicons failed at the sequencing stage. Two of these samples were marinated with either lemon or sweet and spicy marinades, which may either interfere with PCR amplification or result in low amplicon yield which cannot be successfully sequenced. Alternatively, DNA barcode failure in these products may be due to the presence of more than one species, which can co-amplify and produce a mixed electrophorogram.
For the samples which have multiple closely related species, they generated overlapping traces at few specific sites in the electrophorogram which were called as ambiguous bases. Indeed, the light tuna products may very well have contained multiple species, as FDA regulations allow for multiple species to be used in the production of canned light tuna as long as the color of the tuna meat is not darker than Munsell value 5.3 (FDA, 2013a).As for the remaining samples (41 products), the 6 mini-barcode primer sets showed different success rates with each tested product group.

Overall, primer sets SH-B, SH-C, SH-D, and SH-E had significantly higher proportions of sequencing success compared to the full barcode (Z-test, two-sided, P values all. This study presents a DNA mini-barcoding system for species identification applicable to heavily processed fish products. Six mini-barcode primer sets were developed, with one primer set in particular showing a high rate of success for identification of heavily processed products at the species or genus level. The additional primer sets developed showed promise as supplemental tools to be used in cases where the initial primer set fails. All mini-barcode primer sets showed increased performance for species identification in heavily processed products as compared to full-length DNA barcode primers. Additionally, the mini-barcoding system provides a new avenue for the utility of next-generation DNA sequencing for authentication of mixed products that may contain multiple species and experienced different levels of DNA un-friendly commercial procedures.
Overall, the mini-barcode system developed here provides a means to identify species in heavily processed products and may be used for the detection and enforcement of species substitution on the commercial market. 1.Woolfe, M.
& Primrose, S. Food forensics: using DNA technology to combat misdescription and fraud. Trends Biotechnol 22, 222–226 (2004). 2.Marko, P.
Assassins creed origins sun dial puzzle. Fisheries: mislabelling of a depleted reef fish. Nature 430, 309–310 (2004). 3.Hellberg, R. & Morrissey, M. Advances in DNA-based techniques for the detection of seafood species substitution on the commercial market. J Lab Autom 16, 308–321 (2011). 4.Wong, E.
DNA barcoding detects market substitution in North American seafood. Food Res Int 41, 828–837 (2008). 5.Wallace, L. DNA barcodes for everyday life: routine authentication of Natural Health Products. Food Res Int 49, 446–452 (2012). 6.Teletchea, F.
Barcode Software
Molecular identification methods of fish species: reassessment and possible applications. Rev Fish Biol Fisher 19, 265e293 (2009).
From healthcare to education, the uses of barcodes are far-reaching and, as a result, so are the benefits of the software used to create and scan these barcodes. Aren’t using barcode software yet? Here are five key that might just make you change your mind:. Easily Create Quick Custom LabelsBarcode software creation is an obvious but important benefit of barcode software. Wasp’s includes more than 100 pre-built templates, making it easy to create inventory labels, asset tags, address and shipping labels, ID badges, and coupons. That means you can rapidly design custom labels, including barcodes, text, and graphics. Simplify Tracking with Continued Serial Numberslike WaspLabeler +2D allows the user to automatically serialize inventory or asset tags to simplify tracking.
WaspLabeler +2D even automatically remembers the number used previously, so you never have to keep track of where you left off. Eliminate Data Entry ErrorsFast-paced environments can make your business susceptible to errors. Eliminating manual data entry by creating barcodes using barcode software means you will cut down on those frustrating mistakes. Improve Accuracy with Database ConnectivityA barcode software solution, like WaspLabeler +2D, can save you time and money with database connectivity. If your data exists in a spreadsheet, SQL server, or within Quickbooks, you can easily connect WaspLabeler +2D to that database. Easily input data into your label, saving you time and avoiding costly errors.
Improve EfficiencyNot every inventory item can be easily scanned at checkout. Solutions like Wasp’s help improve efficiencies of retail checkout with barcoded product lists for small or bulky items.Wasp Barcode is the leading manufacturer of barcode software and barcode software solutions for small to medium-sized businesses. Watch this short video to learn more about.
Weighing ScalesChoose the counting scale right for you! Scale functionality ranges from simple, stand-alone counting and weighing, in capacities from 6 to 60 pounds, to high precision, multi-platform counting with accumulation or data base capabilities including weight, count, & tare weight memory for up to 1,000 items.Designed with CAS reliability, the EC Series Counting Scale stands alone its field. High accuracy and simple to use.
Optional label or receipt printer. Great for use in manufacturing, parts shops, hardware stores, etc.
Comments are closed.
|
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
- Blog
- Animasi Power Point Bergerak
- Dz09 Mtk6261 Backup Hex
- Dupur Thakurpo Full Movie Download Moviescounter
- Rm 208 Flash File Download
- Monitor Dims And Brightens
- Nexus 2.3.4 Full Torrent
- Cisco Packet Tracer Terbaru 2019
- Sims 3 All Expansions Free Download
- Download Driver Hp Color Laserjet 5550dn
- Download Adobe Bridge Cc 2019 Full Crack
- Download Call Of Duty 1 For Pc
- Free Download Jurassic Park
- Remove Most Visited Chrome
- Yugioh Castle Of Dragon Souls
- Skyrim Become A Dragon Mod
 RSS Feed
RSS Feed
